Sommaire
- When the pharmaceutical chain invents its own standards for its invisible links?
- Transporteurs & Bonnes Pratiques de Distribution en gros des Médicaments (BPDG). Quand chaque kilomètre compte pour la qualité
- L’importance des étapes de nettoyage et désinfection dans la maitrise de la contamination en salle propre
- The Impact of Next Generation Sequencing (NGS) on Quality Control of Pharmaceutical Products
- De la contamination à l'amélioration continue : méthode d’investigation des déviations microbiologiques dans l’industrie pharma
- Economic justification for automation & robotics solutions in pharmaceutical microbiology quality control
- Conception et exploitation d'un laboratoire de microbiologie : l’approche collaborative du GIC A3P Laboratoire de Microbiologie
- Bonnes pratiques d’utilisation d’un Poste de sécurité microbiologique de type II au laboratoire analytique microbiologie
- Waste mapping
The Impact of Next Generation Sequencing (NGS) on Quality Control of Pharmaceutical Products.
Pharmaceutical QC regarding nucleic acids in productions has traditionally relied on targeted, low throughput, or low resolution analytical methods.
For instance, for sequencing of either DNA or RNA, Sanger’s based methods, the highly reliable reference for almost 50 years, relied on already known target sequences, framed by primers and di-deoxynucleotides to generate sequencing products that can be analysed by capillary gel electrophoresis (with limited length). Each target may require a dedicated standard operating procedure.

While these techniques remain important, they often lack the precision needed for increasingly sophisticated products and are limited. NGS, also referred to as High-Throughput Sequencing (HTS), by contrast, enables comprehensive, untargeted characterization of nucleic acids from multiple sources—cells, viruses, plasmids, and microbial communities—with exceptional sensitivity and scalability. Different generations of these technologies succeeded to each other. This “platform technology” is called “agnostic” since it doesn’t require any prior knowledge of a target’s sequence. These characteristics make NGS particularly suited to modern QC challenges where product safety and molecular fidelity are critical. Regulatories are prone to encourage NGS applications in QC of pharmaceutical products.
1. NGS as a Game Changer in QC
1. 1. NGS technologies
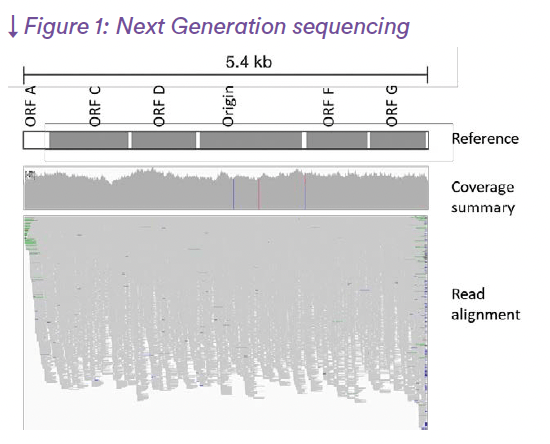
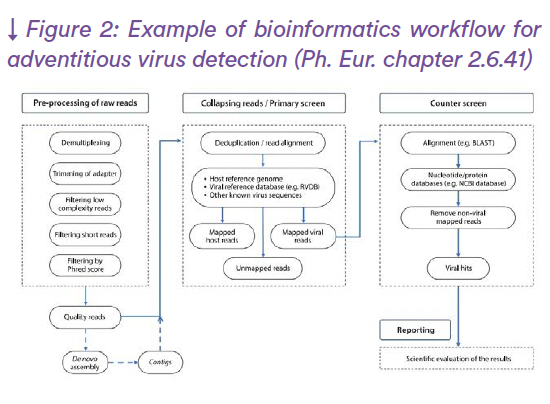
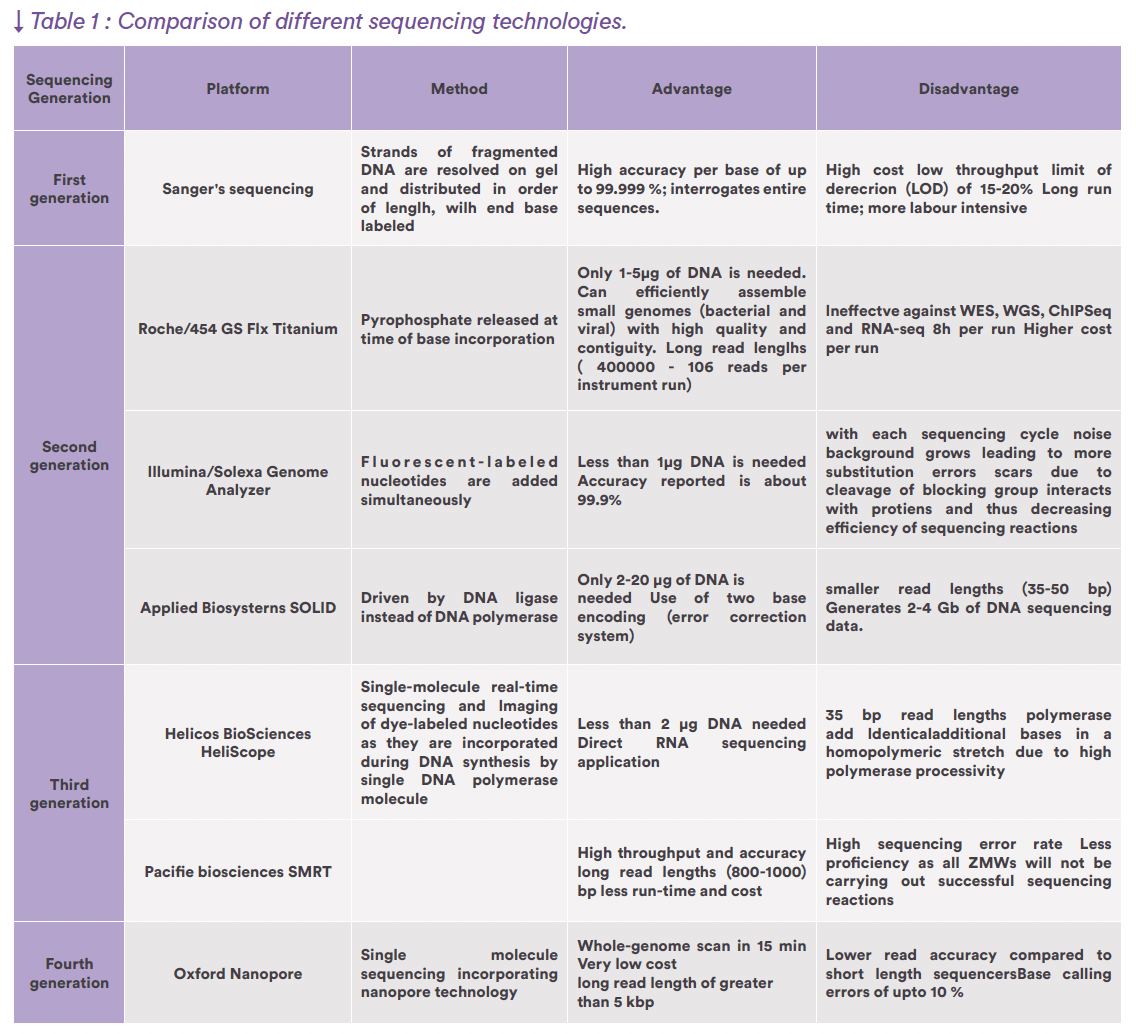
The two main NGS technologies are providing with millions of “Short reads” (roughly in between 120 and 500 nucleotide long sequences) or millions of “Long reads” (roughly in between 800 and 50,000 nucleotide long sequence) (see Figure 1 for Short Read example). They both have pros and cons regarding accuracy or efficiency, while they can be highly complementary. The different generations of NGS technologies are listed in table 1 with their main characteristics (Katara et al., 2024). Bioinformatic treatment of the data would permit their reliable and automatic interpretation using dedicated pipelines of analysis according to the applications (Satam et al., 2023) (see figure 2 for un example of application to adventitious virus detection (from EP chapter 2.6.41)).
Figure 1: Next Generation sequencing: Genome schematic, sequencing coverage, and read alignment for φX174 sequenced by Illumina NextSeq 1000/2000 technology (short read). The linearized φX174 genome (5.4 kb) is shown with major ORFs indicated as grey boxes. Sequencing depth across the genome is displayed in grayscale, followed by individual read alignments illustrating mapping uniformity and coverage decline near genome termini.(Table 1)
(Eurofins BPT Biologics; internal data)
1.2. High Sensitivity and Breadth of Detection.
NGS detects DNA and RNA sequences at extremely low abundance, enabling:
- Sensitive identification of adventitious agents, including bacteria (including mycoplasma), fungi, and viruses—even those that cannot be cultured.
- Genome-wide analysis of contaminants without prior assumptions or specific detection probes (“agnostic” method).
- Simultaneous detection of multiple organisms and impurities in a single assay.
- Sequencing of otherwise hard to sequence sequence Adeno-Associated Virus (AAV) inverted terminal repeats (Namkung et al., 2022) Lentivirus (LT).
This unbiased approach significantly reduces the risk of missing unforeseen contaminants, which is particularly valuable in complex biological matrices.
1.3. Ensuring Genetic Identity and Stability
For biologics, cell banks, plasmids and viral vectors, maintaining genetic fidelity is essential. NGS contributes by:
- Confirming cell line identity through whole genome or targeted sequencing.
- Monitoring genetic drift across manufacturing passages.
- Characterizing vector integrity, including truncations, rearrangements, or plasmid variants.
- Providing full length genome confirmation for gene therapy products.
Compared to Sanger or PCR based assays, NGS provides far deeper insight into sequence heterogeneity and product stability.
1.4. Microbiome and Host Cell DNA Residuals
Regulators expect increasingly precise quantification of process related impurities. NGS supports this by:
- Quantifying host cell DNA fragments with greater sensitivity and size profiling than qPCR.
- Characterizing residual microbial communities in fermentation based processes.
- Enabling metagenomic profiling to assess microbial ecosystems that influence product quality.
For microbial produced APIs, NGS can also help characterize and optimize production strains.
2. Transformational Applications in Biopharmaceutical QC
2.1. Viral Safety Testing
NGS provides a powerful complement or alternative to classical in vitro and in vivo adventitious agent testing (Alston et al., 2025). It can:
- Detect known and unknown viral contaminants.
- Characterize viral genomes in viral vector manufacturing.
- Reduce false negatives associated with culture based tests.
As regulatory acceptance grows (e.g. European Pharmacopoeia, chapter 2.6.41) NGS may streamline release testing for viral vector products.
2.2. Cell and Gene Therapy Manufacturing
Cell therapy materials, CAR T products, and AAV/LV vectors rely on NGS to assess:
- Gene insertion sites.
- On and off target editing for CRISPR based therapies.
- Vector genome integrity.
- Replication competent virus contamination.
These analyses address unique safety concerns inherent to advanced therapies.
2.3. Vaccine Development and QC
NGS supports:
- Strain verification for viral and bacterial vaccines,
- Monitoring attenuation stability in live vaccines.
- Profiling viral quasispecies.
- Characterizing mRNA vaccine templates and integrity.
The COVID 19 pandemic accelerated the adoption of NGS based QC tools in vaccine production.
3. Regulatory Evolution and Acceptance
The regulatory landscape governing the use of Next-Generation Sequencing (NGS), in the viral safety assessment of biopharmaceutical products has evolved markedly over the past decade. International Health Authorities now increasingly recognize NGS based methods as sensitive, broad-spectrum approaches capable of detecting adventitious viral contaminants in cell substrates, vaccines, recombinant proteins, gene therapy vectors, and other biologics.
The World Health Organization (WHO) first acknowledged the potential of high-throughput sequencing in its Technical Report Series for Cell Substrates and vaccines like Yellow fever, Dengue or Polio since 2012, highlighting its possible use as an alternative or complementary approach to existing in vivo and in vitro tests for virus detection.
In parallel, the U.S. Food and Drug Administration (FDA), notably through the PDA/FDA Advanced Virus Detection Technologies Interest Group (AVDTIG) which since became a Working Group (AVDTWG) has supported technological development and scientific dialogue around such advanced sequencing-based viral detection since 2012.
In 2017, the European pharmacopeia in edition 9.3 introduced the use of NGS into two chapters updates, chapter 5.2.3. on Cell Substrates for the Production of Vaccines for Human Use and chapter 2.6.16. tests for Extraneous Agents in Viral Vaccines for Human Use, mentioning the concept of broad molecular methods (such as high throughput sequencing) for broad viral detection: “Tests for viruses using broad molecular methods. In agreement with the competent authority, broad molecular methods (e.g. High Throughput Sequencing) may be used either as an alternative to in vivo tests and specific NAT or as a supplement or alternative to in vitro culture tests based on the risk assessment.”
A significant step was achieved with the publication of the revised ICH Q5A(R2) guideline on viral safety evaluation of biotechnology products in November 2023, which clearly indicates that NGS for adventitious virus detection can be used to replace the compendial in vivo tests, and to supplement or replace the in vitro test , this without head to head comparison between the historical compendial methods and the NGS based test..
The Chinese Pharmacopoeia has similarly integrated NGS into its 2025 Requirements for the Quality Control of Animal Cell Substrates used in the Manufacture of Biologics, positioning NGS as a potential replacement for traditional tests, especially relevant when the sample interferes within the conventional tests.
In Europe, the regulatory framework has drastically progressed with HTS for adventitious virus detection within the creation of a dedicated general chapter, Ph. Eur. chapter 2.6.41 “high-throughput sequencing for the detection of viral extraneous agents”, which will be applicable in April 2026. This chapter provides comprehensive guidance on HTS workflows, methodological design, analytical approaches such as genomics, viromics, and transcriptomics, and detailed expectations for method validation, including the selection of appropriate reference materials and the evaluation of assay performance characteristics.
Finally, in the latest news, the Indian Pharmacopoeia has just announced in its new 2026 edition the publication of a new general chapter the “High-Throughput Sequencing (HTS) based Detection of Viral Extraneous Agents” (new chapter 2.2.35.).
This overview reflects the broad international convergence toward the regulatory acceptance of NGS for adventitious virus detection, especially in replacement of in vivo test, to support viral safety in the production of modern biopharmaceuticals.
Key trends include:
- Encouragement of NGS for adventitious agent detection (e.g. Ph. Eur. Chapter 2.6.41).
- Use in characterization of cell lines and raw materials.
- ntegration into guidelines for gene therapy vectors and mRNA vaccines.
However, regulators emphasize method validation, data integrity, and bioinformatics transparency—areas requiring robust QC frameworks.
4. Challenges and Opportunities
NGS may generate very big volume of data (e.g. Tera Octets of data), it involves a lot of Bioinformatic “pipelines” and resources for assembly of those data, annotation by sequence homology, exclusion of unwanted sequences, comparison of large number of sequences (genes) to databases, etc… Hence, bio-informatician became a more and more important qualification for QC labs members as part of both the challenges and the opportunities.
4.1. Challenges
- Complex validation due to multi step workflows and bioinformatic pipelines.
- Data interpretation variability, requiring stringent standards.
- High computational demands and need for controlled data management.
- Cost and expertise needs, though both are decreasing over time.
4.2. Opportunities
- Replacing or reducing reliance on animal based viral safety tests.
- Real time monitoring of microbial shifts in bioprocesses.
- Enhanced lot to lot comparability using high resolution genomic fingerprints.
- Greater automation and standardized pipelines for QC environments.
- New professions need
5. Conclusion
NGS has emerged as a transformative tool for pharmaceutical QC, offering unprecedented depth, sensitivity, and breadth of analysis. From adventitious agent detection to confirmation of genetic fidelity in complex biologics, NGS strengthens product safety, consistency, and regulatory robustness. As analytical methodologies mature and regulatory frameworks evolve, NGS is expected to become a standard component of QC workflows across biologics, vaccines, cell and gene therapies, and advanced pharmaceutical modalities. New professions will emerge and develop such as those of bioinformaticians for programming and analyses.
References
- 1. Alston et al., Biologicals, Volume 90, April 2025, 1-6 Validation of a Next Generation Sequencing Method for adventitious agents detection in a live vaccine matrix
- 2. European Pharmacopeia; chapter 5.2.3. cell substrates for the production of vaccines for human use, January 2018 and chapter 2.6.16., Tests for extraneous agents in viral vaccines for human use, January 2024.
- 3. European Pharmacopeia; 12.2. chapter 2.6.41, 1-18 “High-throughput sequencing for the detection of viral extraneous agents”, Applicable 04-2026.
- 4. Chinese Pharmacopeia ChP, 2025 edition, chapter on requirements for preparation and quality control of animal cell substrates used for production of biologics.
- 5. ICH Q5A (R2), Viral safety evaluation of biotechnology products, November 2023, chapter 3.2.5.2. and IWG Training on viral safety evaluation of biotechnology products derived from cell lines of human or animal origin, May 2025.
- 6. Katara et al., Journal of Chromatography Open, Volume 5, May 2024, 100-121 “Evolution and applications of Next Generation Sequencing and its intricate relations with chromatographic and spectrometric techniques in modern day sciences”
- 7. Namkung et al., Human Gene Therapy, Volume 33, September 2022, N° 21 and 22, 1187-1196 “Direct ITR-to-ITR Nanopore Sequencing of AAV Vector Genomes”,
- 8. Göke et al ., Nature Methods, Volume 22, April 2025, 801–812 “A systematic benchmark of Nanopore long-read RNA sequencing for transcript-level analysis in human cell lines”
- 9. Satam et al., Biology, Volume 12, 997-1022” Next-Generation Sequencing Technology: current trends and advancements”
- 10. WHO, Technical Report Series on cell substrates, Yellow Fever vaccine, Dengue Vaccine, since 2012
Partager l’article
Thierry BONNEVAY
Arnaud CARLOTTI